Scientists Decode Genome of Pacific White Shrimp in Scientific First
Date:2019-01-23 | 【Print】 【Close】
A research team led by Prof. XIANG Jianhai and Prof. LI Fuhua from the Institute of Oceanology of the Chinese Academy of Sciences (IOCAS), in collaboration with domestic and international colleagues, successfully finished the decoding of the highly complex genome of the Pacific white shrimp Litopenaeus vannamei.
Penaeid shrimp is regarded as the representative species of decapod crustaceans. The Pacific white shrimp Litopenaeus vannamei is a world-wide aquaculture species with its production accounts for more than 2/3 of shrimp aquaculture production.
Environment deterioration, disease problems and germplasm degradation are becoming the most severe threat to the shrimp aquaculture. Understanding their genetic basis to manage the pathogen infection and the genetic improvement is very urgent for shrimp species. However, the whole genome sequencing of shrimp faced a huge challenge due to their big genome size, high repeat sequence and high heterozygosity.
To solve the above technological issues, integrated omics studies, including combination approach of the second and the third generation sequences, transcriptomics, proteomics and bioinformatics, have been implemented. After more than 10 years of efforts, the first shrimp de novo reference genome in the world has been completed. The study was published in Nature Communications.
The L. vannamei genome covers ~1.66 Gb (scaffold N50 605.56 Kb) with 25,596 protein-coding genes. It holds the highest proportion of simple sequence repeats (SSRs, more than 23.93%) among the sequenced animals, presenting a challenge for genome sequencing and assembly.
The high quality genome assembly of L. vannamei provides the opportunity to understand various biological processes of shrimp at the genome level. The most prominent characteristic of the coding region of the shrimp genome is the expansion of a series of genes related to visual system, nerve signal conduction and molting.
These gene expansions might have provided the shrimp with its excellent eyesight and rapid nerve signal conduction, better equipping it to adapt to its benthic habitat. The shrimp genome assembly also sheds light on the molting process and provides important evidences for exploring the similarities and differences between crustaceans and other ecdysozoans, including insects.
As a vital process in shrimps, molting is tightly linked to growth, metamorphosis and reproduction. Elucidating the regulatory mechanisms of molting will provide useful clues for genetic analysis of these important processes. The genomic analysis on the breeding population and wild population of shrimp reveals that the farmer’s breeding practice of only 30 years has already exerted a significant impact on the genome of the L. vannamei broodstocks.
An increase in aquaculture production and efficiency can be achieved through genetic improvement of cultured stocks. The assembled shrimp genome and the large amount of SNP markers will provide a useful resource for the application of genome-wide association studies and genomic selection, and thus accelerate genetic improvements in shrimp culture.
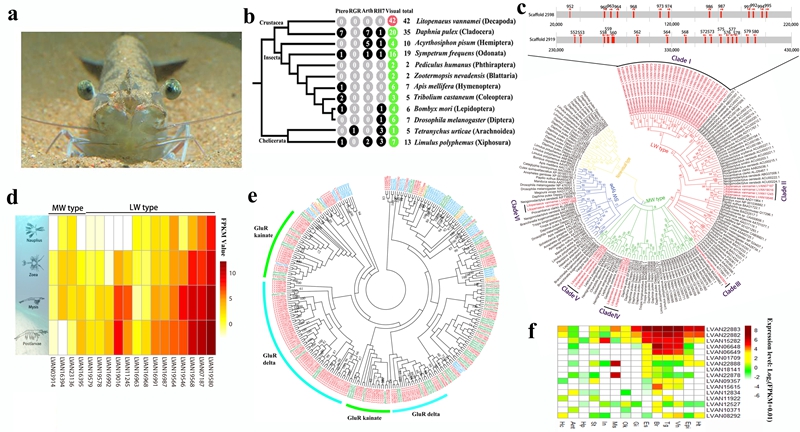
The Opsin and iGluR gene families in the shrimp genome to show its adaptions to benthic environment.
Reference
https://www.nature.com/articles/s41467-018-08197-4
 Address: No. 7, Nanhai Road, Qingdao City Postcode: 266071
Address: No. 7, Nanhai Road, Qingdao City Postcode: 266071